New tool RODEO captures breadth of microbial biosynthetic potential
In an age of booming biotechnology, it might be easy to forget how much we still rely on the bounty of the natural world. Some microbes make us sick, some keep us healthy, while others continue to give us some of our best cures in the form of naturally occurring products such as penicillin and tetracycline.
A new bioinformatics advance from the University of Illinois reveals the power of ‘big data’ genome technology to help us make better use of nature’s inventions: a team of researchers led by Associate Professor of Chemistry Douglas Mitchell has created a tool that searches through microbial genomes, identifying clusters of genes that indicate an organism’s ability to synthesize therapeutically promising molecules.
In a Nature Chemical Biology article (DOI: 10.1038/nchembio.2319), lead authors Jonathan Tietz and Christopher Schwalen and their colleagues in Mitchell’s laboratory describe how their custom software learns to recognize predictive genomic features.
“With genome sequencing going at the pace it has . . . there's a dearth of functional information about what these genes are doing,” Mitchell said. “It becomes increasingly important to make sense of and interpret metabolic pathways, especially biosynthetic gene clusters encoded by microbes.”
Mitchell’s group is particularly interested in a class of molecules commonly referred to as RiPPs, a friendly acronym for a long name: ribosomally synthesized and post-translationally modified peptides. RiPPs, like proteins, are made of chains of linked amino acids, are encoded by genes, and undergo chemical modification (carried out by other proteins) after they are made. Compared with proteins, RiPPs are smaller and the modifications they undergo alter their structure more dramatically.
RiPPs may seem unfamiliar, but they are already present in the average consumer’s daily life. A bacterially-produced RiPP called nisin, for example, has been used as a pathogen-fighting additive in dairy products, meats, and beverages such as beer since the 1960s.
“RiPPs have some particular advantages compared to other, more traditional, classes of natural products. They’re usually larger and more structurally complex,” which allows them to interact with cellular machinery in ways a smaller molecule cannot, Tietz explained. More points of contact with their cellular targets means RiPPs can hang on better and perform more complicated tasks. “At the same time, despite their complexity, RiPP biosynthesis . . . makes for greater potential for genetic re-engineering of natural products to tailor physical and pharmacological properties,” he noted.
For all their advantages, RiPPs present a challenge; it is hard to discover new ones. Traditionally, researchers found potentially useful natural products by screening microbes based on their biological activity. After decades of such efforts, which revealed a range of products including some RiPPs, the low-hanging fruit has been plucked; searches turn up the same common compounds over and over again. Mitchell and colleagues are members of a multi-laboratory research group at the Carl R. Woese Institute for Genomic Biology (IGB) that has found a way to “climb higher” and uncover novel natural products: genome mining.
The Mining Microbial Genomes research theme at the IGB aims to speed drug discovery by searching through the genomes of microbes, essentially skimming through cells’ recipe books to see what they might be able to produce, before actually persuading them to do so in a laboratory setting. In this way, researchers can greatly increase the odds that they will isolate a compound that has never been seen before. However, this method relies on the ability to predict what a group of genes might be capable of producing.
“In a practical sense the question became, is there a better way to harness available genomes for augmenting these discovery pipelines,” said Schwalen. “That's where we started.”
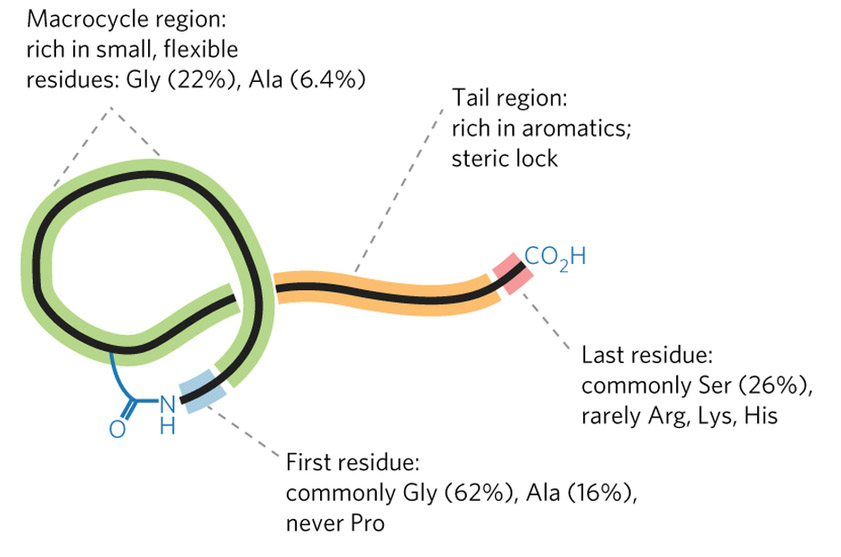
Mitchell’s group faced a tough challenge: creating software that could recognize the groups of genes whose products work together to synthesize a RiPP. They decided to make it even tougher by focusing on a class of RiPPs called lasso peptides, named for their looping structure. The clusters of genes that produce lasso peptides are small and generic-looking, making them difficult to identify even in a manual search.
“If you want to show that you have a useful tool, you pick the hardest example,” Mitchell said. “But also, as a chemist, lasso peptides are extremely interesting. Peptides that are used as drugs cannot be given orally” because they would be digested, he explained. “Lasso peptides are different. You can actually boil these, you can throw proteases at them, you can autoclave them and they don't lose their activity, they are basically a little peptide knot that is extremely resistant to such assaults.”
The informatics tool that Mitchell’s laboratory designed, named RODEO (Rapid Open reading frame Description and Evaluation Online), dealt with the lasso challenge in part through a machine learning approach. They trained the software on known examples of lasso-producing gene clusters, allowing the program to hone in on key features. The resulting software robustly identified promising gene clusters in a broad array of microbial genomes, and could be easily customized to search for the gene clusters of other classes of RiPPs as well.
RODEO identified 1300 novel lasso peptides, including several with particularly unusual structures that make them promising as potential therapeutics; the researchers confirmed that the empirically determined structures matched those predicted by the software.
“We can now use genomic prioritization to find molecules that without any doubt are structurally novel,” said Mitchell. "The challenge is, is that a useful molecule or not? But the more molecules you can connect to genes, the better informed we're going to get. So that's the next 10 years of discovery.”
This work was supported by the National Institutes of Health, the American Chemical Society, the David and Lucile Packard Foundation, and Robert C. and Carolyn J. Springborn Endowment for Student Support Program.
